Apache Spark and Essentia are both scalable distributed processing platforms for big data workloads. They both allow to quickly transform data at scales larger than Python can handle effectively. Apache Spark is supported in Amazon EMR, and Essentia is offered through AWS Marketplace.
Performance differences across platforms are not well-known, so I tested both to learn how they perform comparing speed differences across the entire process from dataset transformation to exporting results to S3.
Goal
Compare performance creating a pivot table from Twitter data already preprocessed like the dataset below and export via a compressed csv format into Amazon S3 by
- PySpark on EMR
- Essentia
The transformed data is sparse so compression efficiently reduces the size of the final result.
Before:
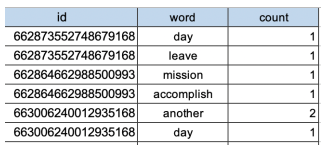
After: [After pivoting with key: id label: word value: sum of count ]
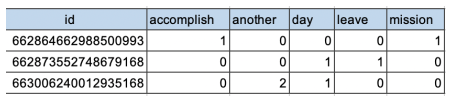
Environments
PySpark on Amazon EMR
-
- Instance Type: m4.large (4 vCore, 8GiB memory, 2vCPU)
- Number of Instances : 1 (master node only)
- Hadoop version: 2.8.5-amzn-4
Pyspark: Python version 3.6.8
mapreduce.fileoutputcommitter.algorithm.version: 2 (This is the default value since Spark 2.0.1. It enables to avoid temporary writes to S3. To know more about this setting, please read XXX)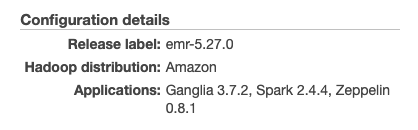
Essentia
-
- Instance Type: m4.large (4 vCore, 8GiB memory, 2vCPU)
- Number of Instances : 1 (master node only)
- Essentia : 4.0.0.2
AQ Tools : AQ Tools : 2.0.1-2 — Wed Aug 14 17:00:00 PDT 201
Data
A. id-word-count data from Twitter
-
- Already cleaned and finished text processing
- Picked top 20,000 words in terms of frequency of occurrence
- 1,696,011 rows x 3 columns
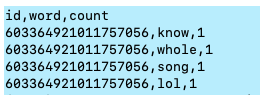
B. Size: 43.8MB
Test Result
PySpark on Amazon EMR
-
- Total Runtime: Couldn’t complete the jobs due to insufficient memory
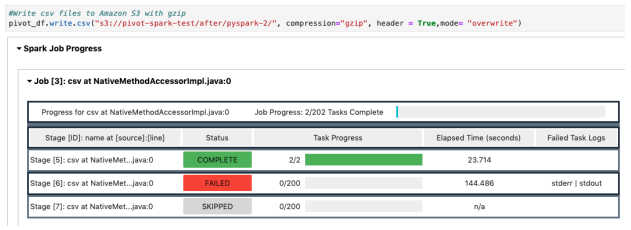
-
- Error Log:
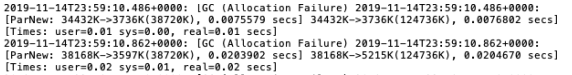

Essentia
-
- Total Runtime: 120 secs (Average runtime from 3 times of the execution, 120,119 and 122 secs)
Pivot execution and export data into S3: 120 secs - Output result:
Compressed size: 21.7MB / Decompressed size: 12.4GB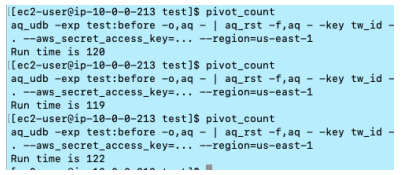
※ For reference the scripts used are placed in the Appendix at the end of this blog.
- Total Runtime: 120 secs (Average runtime from 3 times of the execution, 120,119 and 122 secs)
In the results shown above, Spark couldn’t finish the jobs on m4.large instance with 8 GiB memory while Essentia finished in 2 minutes. Essentia doesn’t use as much memory when creating a pivot table and saving files in S3, instead streaming results directly to S3. This enables saving while compressing. In fact, memory usage during the process didn’t exceed 10% until the job was done. On the other hand, Spark needs memory allocation during execution.
A second test added one more instance as a worker node to both Spark and Essentia, and we evaluated performance again.
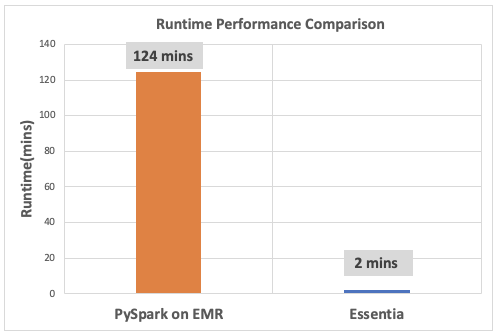
PySpark on Amazon EMR
a. Directly save compressed csv in S3
Total Runtime: 124 mins 21 secs
-
-
- Assign pivot: 21 secs
- Export data to S3: 124 mins
-
b. Save compressed csv in HDFS and copy to S3
Total Runtime:
-
-
- Assign pivot: 21 secs
- Export data to HDFS: 125 mins
- Copy to S3: 1 mins 49 secs
-
Essentia
a. Total Runtime: 119 secs
Pivot + Export data to S3
In the chart above we see that PySpark was able to successfully complete the operation, but performance was about 60x slower in comparison to Essentia.
Appendix
1-a. Performance Notes of Additional Test (Save in S3/Spark on EMR)
- Assign pivot transformation
- Pivot execution and save compressed csv to S3
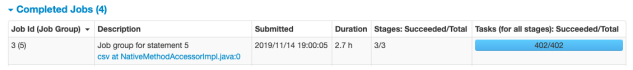
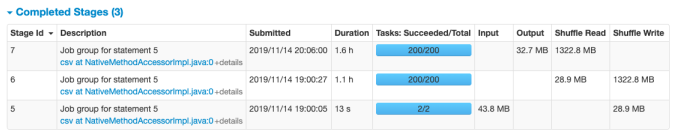
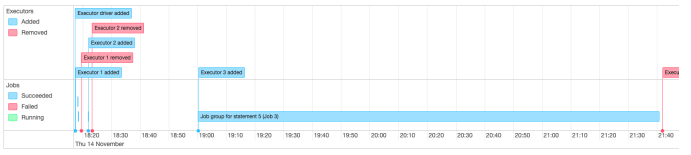
1-b. Performance Notes of Additional Test (Save in HDFS/Spark on EMR)
- Assign pivot transformation
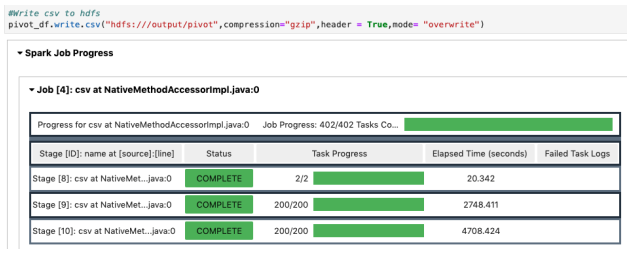
- Pivot execution and save compressed csv to S3
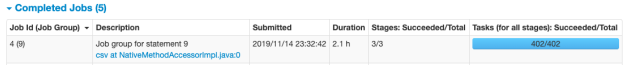
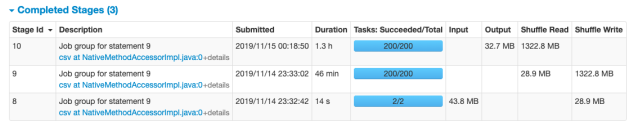

2. Performance Notes of Additional Test (Essentia)


3. Scripts
- PySpark
%%info
#Import SQLContext
from pyspark.sql import SQLContext
#sc is automatically created when initialize notebook on EMR
sql = SQLContext(sc)
df_top_only = (sql.read
.format("com.databricks.spark.csv")
.option("header", "true").option("inferSchema", "true")
.load("s3://pivot-spark-test/before/only_top_words/twitter_word_top20000.csv"))
spark.conf.set("spark.sql.pivotMaxValues", 100000)
#Assign pivot transformation
pivot_df = df_top_only.groupBy("tw_id").pivot("word").sum("count").fillna(0)
#A)Execute pivot transformation and write csv to Amazon S3 with gzip
pivot_df.write.csv("s3://pivot-spark-test/after/pyspark/", compression="gzip", header = True,mode= "overwrite")
#B)Execute pivot transformation and write csv to hdfs with gzip
pivot_df.write.csv("hdfs:///output/pivot", compression="gzip",header = True,mode= "overwrite")
- Essentia
#!/bin/bash
OUTPUT="--debug --s3out pivot-spark-test:/after"
COL="s:id s:word i:count"
DB_SPEC="s,pkey:id s,+key:word i,+add:count"
OPT="--debug"
#Set the input data path in S3
ess select s3://pivot-spark-test
ess category add twitter "before/only_top_words/twitter_word_top20000.csv" --noprobe --overwrite
#Create database
ess server reset
ess create database test --port 0
ess create table before $DB_SPEC
ess server commit
#Import data into database
ess stream twitter "*" "*" "aq_pp -f,+1 - -d $COL -imp test:before" $OPT
pivot_count() {
start_time=`date +%s`
#Pivot and Export the compressed file to s3
ess exec "aq_udb -exp test:before -o,aq - | aq_rst -f,aq - -key id -lab word -val count -o - | gzip" --master $OUTPUT/after_pivot_with_ess.csv.gz
end_time=`date +%s`
run_time=$((end_time - start_time))
echo "Run time is $run_time"
}
#Execute function
pivot_count
